FAQs
Scan Alignment
How long does a typical model-fit to scan data (500k-1000k vertices, covering most of the scanned body) take, if one would perform the calculations on a normal offline PC (let’s say i7, 32GB Ram)?
The processing for meshcapade.me is online, so your PC setup is not relevant. However, we have created a downloadable version of the software which can run on a PC. For a PC you described (i7, 32GB Ram) the runtime will be between 15-25 minutes. If you would like to find out details of the downloadable software, we can share the licensing details with you.
Do you recommend specific hardware like graphic cards to speed up the process?
Not at the moment. We are working on GPU-enabled version of the software which will allow much faster processing. We will announce this later this year.
Do you have different “fit-stages” where the parametric model is generated first and other fine-fit procedures are added consecutively?
Yes, as you might have already seen on meshcapade.me, when you upload a 3D scan, there is one option called “Refinement”. This option allows you to set whether the additional refinement stages are allowed to be run. If this is set to
- “none” then the output will be directly from the SMPL model’s shape space. Best to use when the input scan is extremely sparse or noisy (e.g. scan have multiple layers of point-cloud surfaces).
- “low”: it will run a refinement stage which allow the vertices for the output mesh to move more freely away from the SMPL model shape parameters. This is best to use if the input 3D scan is noisy (e.g. scans from multi-sensor systems, but also containing spurious scan points that are not expected in our model, like hair, clothing etc).
- “standard”: allows higher refinement. Use this setting when input 3D scan is not too noisy (e.g. high resolution 3D scanning systems).
- “high”: highest refinement level. Use this setting when input 3D scan is extremely clean with no spurious noise (e.g. if the scan has been cleaned up through a separate process).
My Scan alignment doesn’t look right, what went wrong?
Problems with Internal Geometry
The scan alignment process is designed to work with 3D meshes acquired through 3D scanning process.
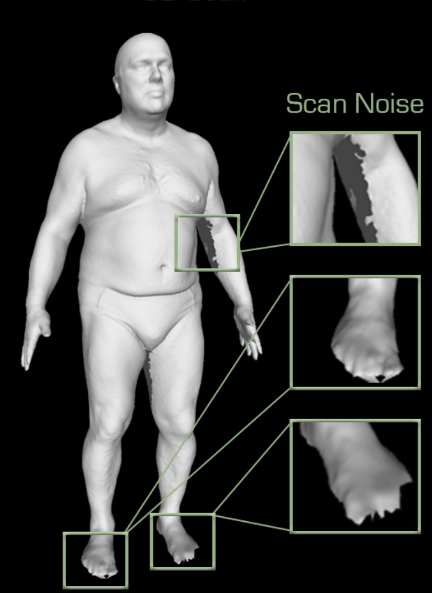
Scans from a 3D scanner can be noisy, this noise is expected and our scan alignment process can handle these.
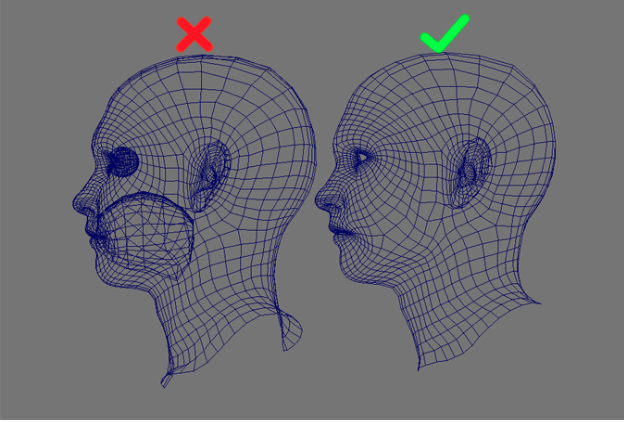
However, the 3D scans are expected to only contain an outer surface of the body, and no internal surfaces. Generally when artists create 3D body models in some 3D animation program, they might create additional geometry on the inside of the body surface, e.g. An artist might create internal geometry for eyes and mouth for the face, as shown below:
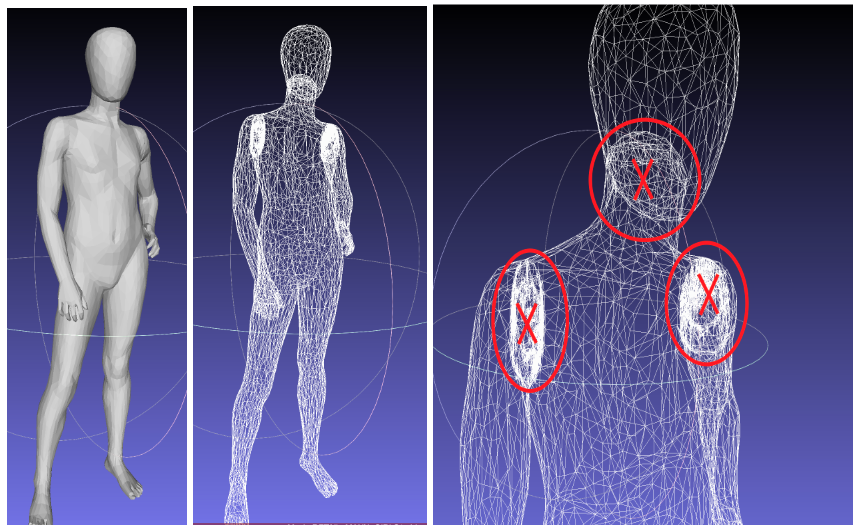
Or there the limbs and neck joints might be created as separate geometry objects, so they will have additional geometry where the limb closes off in the 3D model. These internal geometry pieces have to be removed before they can be sent for processing on meshcapade.me.
meshcapade.me API
What exactly is the API?
The API is a digital human cloud platform for creating animation-ready avatars from scans, hand-measurements, and more. You can find out more details about it here.
How can I license the API?
We offer different pricing options and packages for the API. For more information, please check our licensing page here. If you have higher volumes of data, please contact us on info@meshcapade.com.
What am I allowed to do with the generated avatars? Can I share it with my users?
The output files are free for all uses under SMPL Body license, please contact us on info@meshcapade.com.
How is security being handled for the API?
You can review our Privacy Policy here.
Is your API available as an offline solution/SDK?
Yes. Contact us at info@meshcapade.com.
Is your API available on a mobile application?
Yes. Contact us at info@meshcapade.com.
Is there a trial for the API?
When you choose the pay as you go option, you receive 10 Free credits on initial sign up that you can use to try out the API.
Is there a sample output for the API?
Yes, we have some sample outputs for our bodies-from-scans and bodies-from-measurements API. Please contact us on info@meshcapade.com to share these sample files with you.
MOCAP Processing: IMU Data
How does MoSH work with IMU data?
Short answer: IMU mocap isn’t ideal but if you have the 3D marker trajectories from the IMU system, how about you guys send us a sample of the data and we’ll test how well it works.
Long answer: MoSH is designed to work with optical mocap. It uses the 3D location of the optical markers to fit the SMPL model into each frame of data. The problem with IMU based mocap systems is that you might get the 3D locations of the markers, but there is a general drift associated with IMUs that can cause huge errors over time. Therefore, usually the best option to fit SMPL to IMU is to use any of the IMU based works (e.g. deep initial poser - this is a bit old, but I’m sure there are newer methods out there as well). At Meshcapade we don’t have a license for any of these IMU based works though since most are exclusively for academic use only. But we can still try to use Mosh with certain stronger constraints to make it work in spite of the drift and noise in the 3D marker locations from IMU. But we can’t guarantee how well it will work but happy to give it a try.
Here is a sample file that shows the format for marker data that the mosh code expects:
SMPL - Technical Details
I want to use AMASS but I only want SMPL parameters, not SMPLH. How do I get that?
SMPL-H is exactly SMPL with the addition of an articulated hand parameters. Hand parameteres can be ignored by ignoring the joints 22 and above. So the remaining parameters, ignoring the hand, are exactly the same as SMPL (ie betas and thetas.
What is PA-MPJPE? What is a Procrustes Problem?
MPJPE stands for Mean Per Joint Position Error. Per joint position error is the euclidean distance between ground truth and prediction for a joint. MPJPE measures the mean of per joint position errors for all joints. Most methods estimate the 3D pose and shape in the camera coordinate frame not in the world. Often, the bodies are tilted relative to the world, and sometimes flipped. This results in high errors even if the pose is roughly right. MPJPE is calculated after translating the root (‘pelvis’) of estimated body to the groundtruth root. But, this does not get rid of the rotation issues.
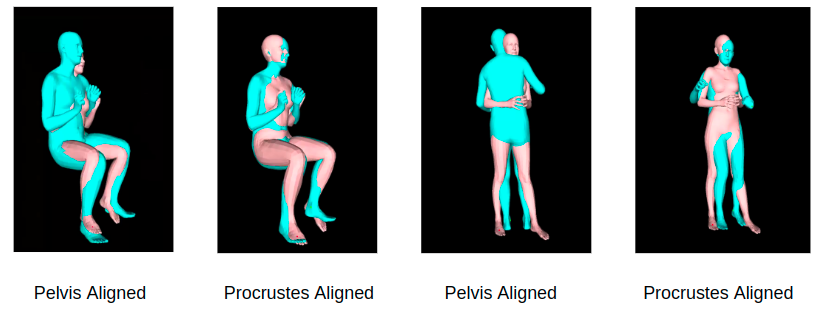
PA-MPJPE uses Procrustes alignment (PA) to solve for translation, scale and rotation between the estimated body and the ground truth. MPJPE is calculated after the estimated body is aligned to the ground truth by the Procrustes method, which is a similarity transformation.
Procrustes Problem: Given correspondences of points \(A_i \in \mathbb{R}^3\) and \(B_i \in \mathbb{R}^3\) find the scaling, rotation, and traslation transformation, called similitude transformation that satisfies
\[A_i = sRB_i + T\]for \(R \in SO(3), T \in \mathbb{R}, and s \in \mathbb{R}^+\).
How do we solve for R, T from \(n\) point correspondences?
Three non-collinear points suffice: each triangle \(A_{i=1...3}\) and \(B_{i=1..3}\) make an orthogonal basis.
\[(A_{21} (A_{21} X A_{31}) X A_{21} A_{21} X A_{31})\]and
\[(B_{21} (B_{21} X B_{31}) X B_{21} B_{21} X B_{31})\]Rotation between two orthogonal bases (one for the B and one for the A) is unique.
We then solve a minimization problem for \(N > 3\) point correspondences which is the translation is centroid of A points (\(\textbf{x}_i\)) and centroid if B points (\(\textbf{y}_i\)) rotated.
We compute the \(d X d\) covariance matrix
\[S = XWY^T\]where X and Y are the \(d X n\) matrices that have (\(\textbf{x}_i\)) and (\(\textbf{y}_i\)) as their columns, respectively and \(W = diag(w_1, w_2,..., w_n)\).
We finally compute the singular value decomposition, \(S = U\Sigma V^T\), to get the rotation and optimal translation.
Please refer to this document for more detailed derivation.
Sample code for such transformation is available here.
How do I add soft-tissue dynamics to SMPL?
You can animate soft-tissue dynamics with DMPL parameters. AMASS uses 8 DMPL parameters to extract soft-tissue motions realistically from a sparse set of markers.
from human_body_prior.body_model.body_model import BodyModel
bm_fname = osp.join(support_dir, 'body_models/smplh/{}/model.npz'.format(subject_gender))
dmpl_fname = osp.join(support_dir, 'body_models/dmpls/{}/model.npz'.format(subject_gender))
num_betas = 16 # number of body parameters
num_dmpls = 8 # number of DMPL parameters
bm = BodyModel(bm_fname=bm_fname, num_betas=num_betas, num_dmpls=num_dmpls, dmpl_fname=dmpl_fname).to(comp_device)
faces = c2c(bm.f)
Please refer to this notebook from the AMASS repository for visualization of DMPL bodies.
With (left) and without (right) DMPL parameters:
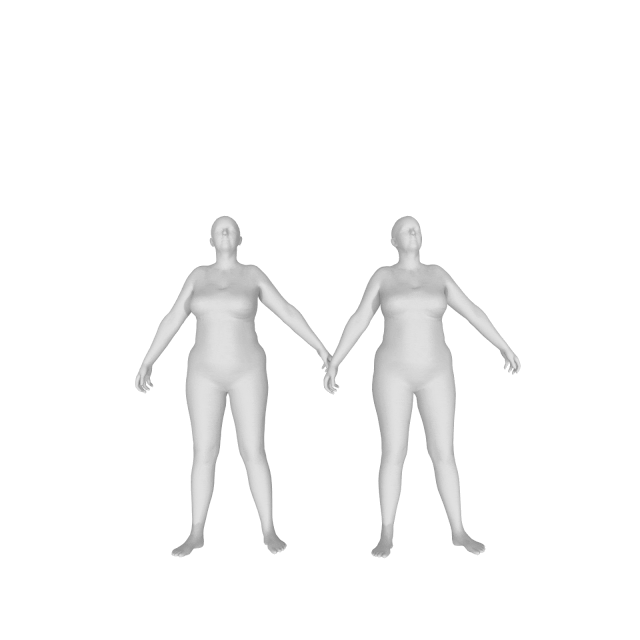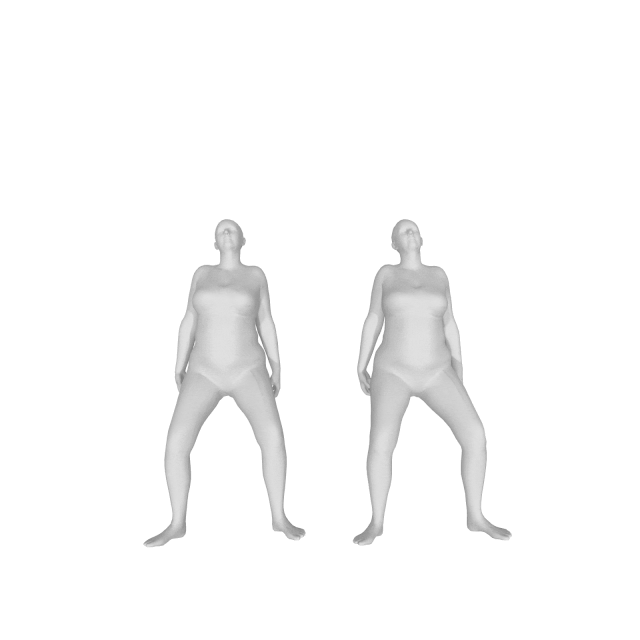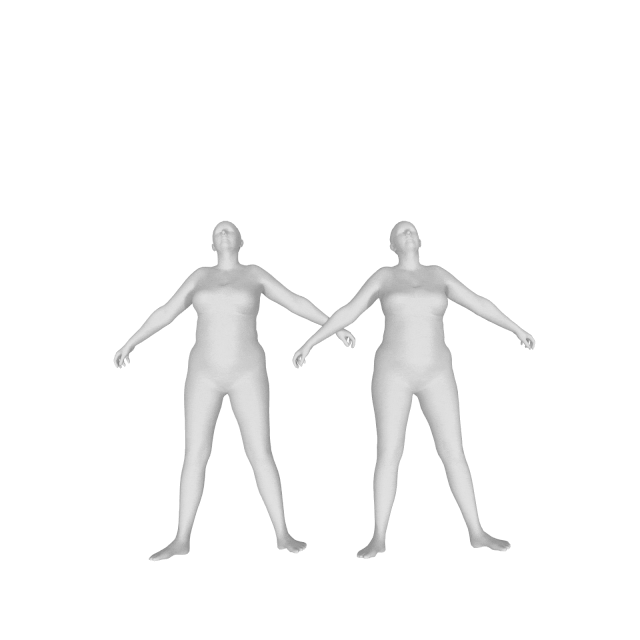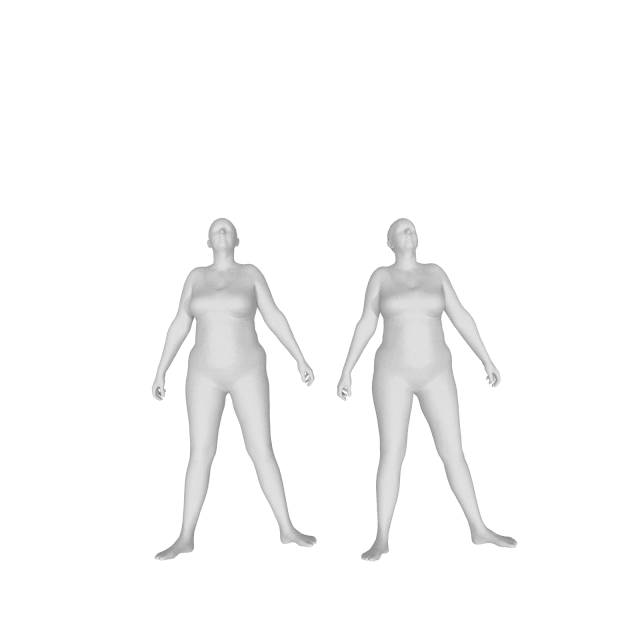
How do I get a gender neutral model?
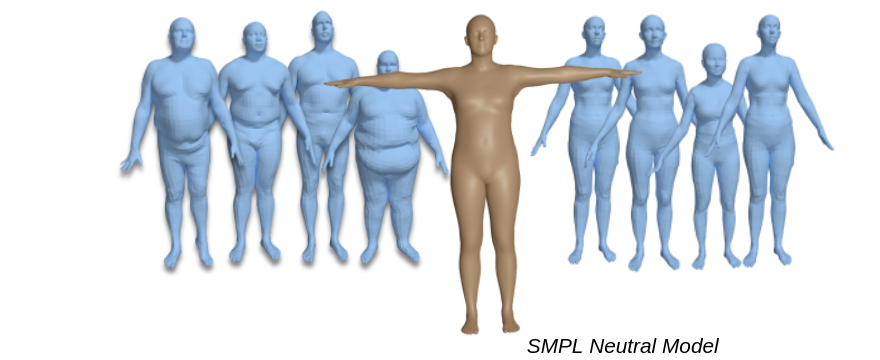
SMPL gender neutral models are available for SMPL , SMPL+H , SMPL-X and STAR. They are created by training both male and female subjects together.
How do I convert between SMPL and SMPL-X (and STAR)?
Please use transfer_model for this.
How do I get SMPL with 300 shape components?
SMPL, SMPLX and STAR are currently available with “full shape space” of 300 shape components. Contact us for licensing.
How do I get just the face (or hand) vertices in SMPL-X?
We provide FLAME and MANO vertex indices of the SMPL-X body as vertex index lists to download. Indexing the SMPL-X body with these index lists returns the vertices that correspond to the MANO and FLAME body parts. This can be done in python as follows:
import pickle
import numpy as np
from psbody.mesh import Mesh
# Load SMPL-X mesh
SMPLX_mesh = Mesh(filename='./smplx_template.obj')
# Load FLAME vertex ids
SMPLX_FLAME_vertex_ids = np.load('./SMPLX_FLAME_vertex_ids.npy')
# Load MANO vertex ids
SMPLX_MANO_vertex_ids = pickle.load('./MANO_SMPLX_vertex_ids.pkl', 'r')
# Extract FLAME vertices from SMPLX_mesh
verts_FLAME = SMPLX_mesh.v[SMPLX_FLAME_vertex_ids]
# Extract MANO vertices from SMPLX_mesh
verts_MANO_left = SMPLX_mesh.v[SMPLX_MANO_vertex_ids['left_hand']]
verts_MANO_right = SMPLX_mesh.v[SMPLX_MANO_vertex_ids['right_hand']]
How do I get the joints out of SMPL?
The easiest way to get joints from SMPL is to use the SMPL class in body_model.py from our SMPLX package.
# Initialize the SMPL constructor object with the correct model path.
smpl_object = SMPL(model_dir='<path_to_SMPL_pkl>')
# Run forward function
smpl_output = smpl_object.forward(betas, pose, global_orientation)
# Extract joints and vertices
joints = smpl_output.joints
vertices = smpl_output.vertices
How do I visualize SMPL in Blender?
Blender accepts .obj and .fbx file types, so once you have the vertices, export them as a mesh and open them in blender. We also offer a blender plugin for more sophisticated usecases.
import trimesh
mesh = trimesh.Trimesh(vertices=vertices,
faces=smpl_object.faces,
process=False,
maintain_order=True)
mesh_fname = 'my_mesh.obj'
mesh.export(mesh_fname)
How do I sample body shapes?
As mentioned, shape space of SMPL is defined by PCA (Principal Component Analysis). PCA returns vectors \(B_i\) of unit norm. For convenience we scale them by \(\sigma\). So to sample according to the Gaussian over body shapes, you can just sample a Gaussian where the covariance matrix is the identity.
To sample a shape where 95% of training subjects lie ([\(-2\sigma, 2 \sigma\)]) in python,
betas_nb = len(model.betas)
amplitude = 2
model.betas[:] = (np.random.rand(beta_nb) - 0.5) * amplitude
How do I keep plausible shapes while optimizing for shape parameters?
You can use shape prior by regularizing the shape vector. This can be done by scaling each dimention by its corresponding variance, i.e.
\[L(\beta) = \sum_i \sigma_i^2 \beta_i^2\]Note that in practice L2 loss is applied on betas directly.
I need a pose prior for my application, how do I do that?
Do check out VPoser. Vposer
- SMPL body pose prior as latent code of variational autoencoder
- Trained on AMASS
- End-to-end differentiable
- Provides a way to penalize impossible poses while admitting valid ones
- Effectively models correlations among the joints of the body
- Can be used to generate valid 3D human poses for data-dependent tasks
- Enables inverse kinematic in batch mode without requiring initialization
I ran SMPLify and the body shape is not right. What happened?
This is natural; estimating a full-3D shape from only a sparse set of 2D joints is highly ambiguous. Think of a pregnant woman; her shape changes drastically, while her skeletal joints stay nearly the same.
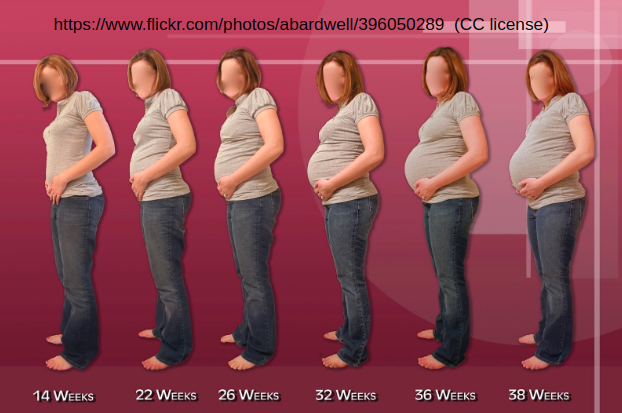
To estimate 3D shape, we need information beyond 2D joints. SMPL is limited to the distribution of the training data. It can’t represent babies, body builders, amputees, and so on.
Can I get the SMPL training data?
Sorry, no.
The SMPL shape space is trained from CAESAR, which we cannot distribute. STAR uses CAESAR and SizeUSA and the same applies. Our pose dataset is also not available due to human subjects limitations. But the FAUST dataset does provide registered poses.
How can I train my own SMPL model?
Currently we do not have training code online.
Good (well registered) training data is key. This is very hard to produce.
It is not so easy to train a SMPL model from scratch.
Curating the data, evaluating intermediate models, fixing problems, adding more data, etc. is all necessary to avoid spurious long-range correlations and artifacts.
How does the UV Map work? Do you have UV maps for SMPL, SMPL-X, etc?
UV maps provide a mapping of each 3D mesh vertex into 2D image space by unwrapping the 3D mesh surface. This technique allows then to map image color information onto the 3D model at a high quality even on low-resolution meshes.
- UV maps for SMPL and SMPL-X are provided on the SMPL/SMPL-X websites.
- Blender SMPL-X add-on (github page) already has UV maps setup.
- UV maps are identical between male/female/neutral models so you can easily swap textures.
Blender SMPL Plugin Questions
Where can I get latest blender plugin?
It is available on our public github page.
I want to pose SMPL myself. How can I do it?
Please use our blender plugin for this. Use our blender plugin to
- bring SMPL/SMPLX model into Blender.
- pose SMPL/SMPLX models.
- export current full-body pose in Rodrigues format to console for later use in Python code.
- auto-calculate pose corrective weights for current pose.
How do I generate animations in FBX format?
- Use the SMPL-X for Blender add-on
- Keyframe the desired motions with Blender pose tools
- For each individual frame keyframe pose corrective weights using the SMPL-X Blender add-on functionality. This can be automated with Blender Python API. Future versions of the SMPL-X Blender add-on will help automating this process.
- Export to FBX.
 Meshcapade Wiki
Meshcapade Wiki